Regularization and Cross-Validation
TRF.train(...) supports both direct fixed-ridge fitting and cross-validated
regularization search.
Scalar Ridge
Pass one scalar to fit directly:
model.train(
stimulus=stimulus,
response=response,
fs=fs,
tmin=0.0,
tmax=0.120,
regularization=1e-3,
)
Use this when:
- you already know a sensible ridge value from previous experiments
- you want the fastest possible fit
- you are running a reproducible pipeline with fixed hyperparameters
In this mode, train(...) returns None because no candidate grid is
evaluated.
Cross-Validated Search
Pass a 1D grid to evaluate multiple candidates:
scores = model.train(
stimulus=stimulus,
response=response,
fs=fs,
tmin=0.0,
tmax=0.120,
regularization=np.logspace(-6, 0, 7),
k=4,
segment_duration=1.024,
overlap=0.5,
)
In this mode:
- one score is computed per candidate regularization value
- the best candidate is chosen automatically
- the final model is refit on all supplied trials using that best candidate
- the candidate grid is stored in
model.regularization_candidates
Internally, ffTRF builds the trial spectra once, then reuses them across
folds and regularization candidates. During validation scoring it also caches
predictor FFTs within each fold, so repeated candidate evaluation avoids
rebuilding the same prediction-side transforms. This applies to both scalar
ridge and banded regularization searches and does not change the selected
solution or the returned scores.
The Meaning of average
The average argument controls how scores are reduced across outputs:
average=True: return one score per regularization candidateaverage=False: keep one score per outputaverage=[...]: average only over the listed outputs
This matters when different output channels behave differently. For example, you might want to select regularization based only on a subset of channels.
The Meaning of k
k="loo"ork=-1: leave-one-out over trialsk=4,k=5, ...: split trials into that many folds
Use leave-one-out when trial count is small and you want maximal use of the data per fold. Use a smaller number of folds when trial count is large and runtime matters more.
Banded Regularization
If your predictor contains grouped features, provide bands so each group can
receive its own ridge coefficient:
model.train(
stimulus=stimulus,
response=response,
fs=fs,
tmin=0.0,
tmax=0.120,
regularization=np.logspace(-5, 0, 5),
bands=[1, 16],
k=4,
)
Example interpretation of bands=[1, 16]:
- first feature belongs to group 1
- next 16 features belong to group 2
In banded mode, ffTRF expands the chosen coefficients into a per-feature
penalty vector internally. Cross-validation still reuses the same cached fold
spectra and validation predictor FFTs, so the banded search stays much cheaper
than rebuilding the full prediction path from scratch for every candidate.
The example below shows how grouped predictors can produce visibly different kernel structure when each block receives its own regularization strength:
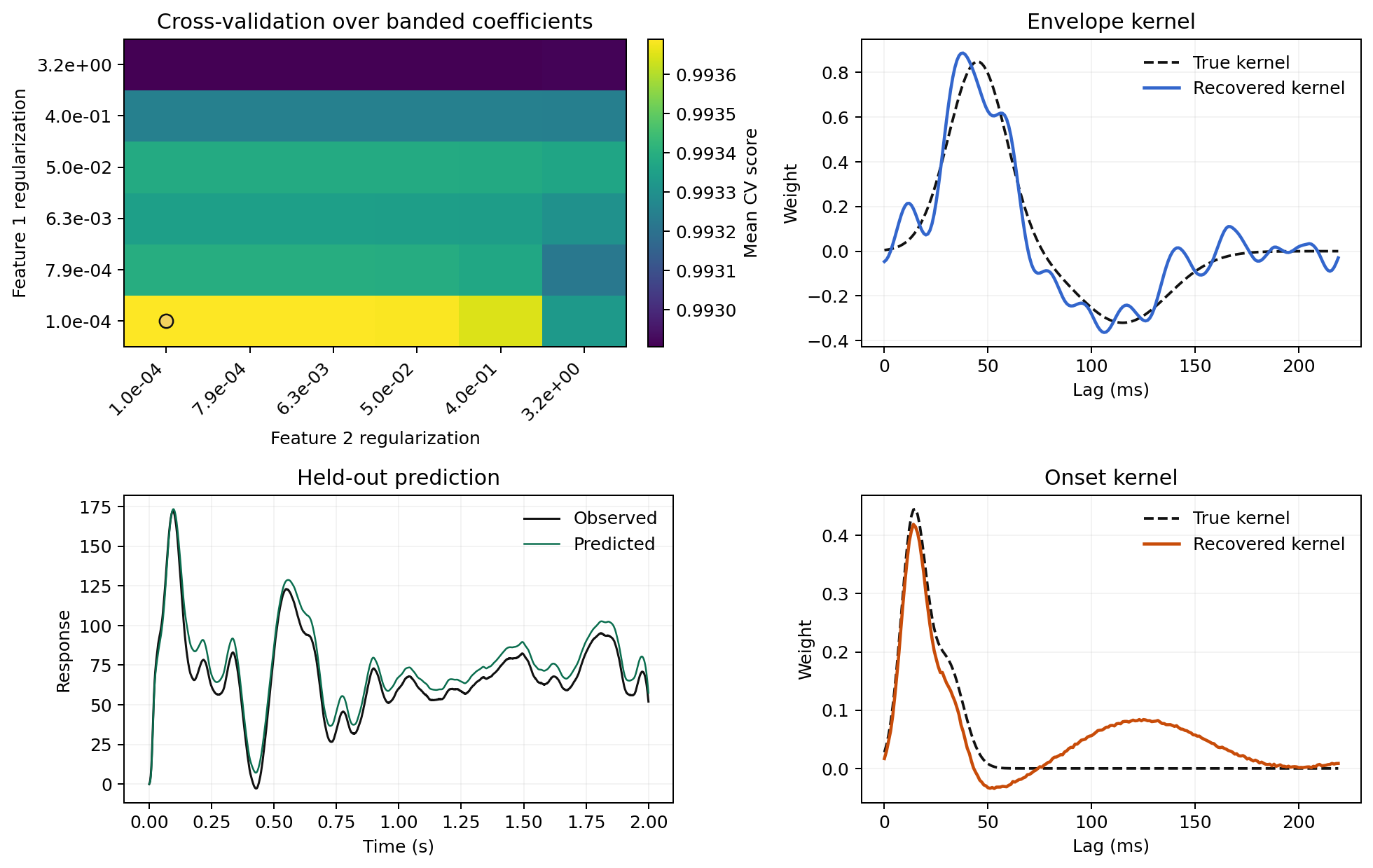
Segment Choices Matter Too
Regularization is not the only stability control. Segment settings matter as well:
- longer segments improve frequency resolution
- shorter segments can increase the number of independent observations
- overlap can stabilize estimates when segments are short
- windowing can reduce spectral leakage in the standard estimator
If a model feels unstable, consider segment settings alongside ridge values. The dedicated Choosing Segment Settings guide collects the practical rules of thumb in one place.
Practical Advice
- Use direct fitting when you already know a sensible ridge value.
- Use
k="loo"when you have only a small number of trials. - Use
k=4ork=5when trial count is larger and runtime matters more. - Use longer segments when you care about lag resolution and narrower spectral smoothing.
- Start with a broad log-spaced grid, then narrow it once you know the useful range for your data.