Getting Started
This page walks through the shortest path from install to a fitted model, while also explaining what the key arguments do and what is stored on the estimator after fitting.
Install
Pixi is the most complete setup for contributors:
pixi install
pixi run import-check
If you only want the package:
pip install -e .
If you want plotting, tests, comparison scripts, or docs tooling in a plain
pip workflow, install the matching extras:
pip install -e ".[compare]"
pip install -e ".[test]"
pip install -e ".[docs]"
For documentation builds, the locked Pixi environment is the most reproducible option:
pixi run -e docs docs-build
For an existing Pixi project, you can link ffTRF directly from GitHub via
Pixi's pypi-dependencies:
[pypi-dependencies]
fftrf = { git = "https://github.com/weigla/ffTRF" }
Then run pixi install. If you want to pin a specific revision, add
rev = "<commit>" to the dependency entry.
First Model
import numpy as np
from fftrf import TRF, inverse_variance_weights
def simulate_trial(
rng: np.random.Generator,
*,
n_samples: int,
kernel: np.ndarray,
noise_scale: float,
) -> tuple[np.ndarray, np.ndarray]:
stimulus = rng.standard_normal((n_samples, 1))
response = np.convolve(stimulus[:, 0], kernel, mode="full")[:n_samples]
response += noise_scale * rng.standard_normal(n_samples)
return stimulus, response[:, np.newaxis]
rng = np.random.default_rng(0)
fs = 512
kernel = np.zeros(60)
kernel[6] = 1.0
kernel[18] = -0.4
kernel[32] = 0.2
trials = [simulate_trial(rng, n_samples=4_096, kernel=kernel, noise_scale=0.05) for _ in range(6)]
stimulus = [trial_stimulus for trial_stimulus, _ in trials]
response = [trial_response for _, trial_response in trials]
model = TRF(direction=1)
cv_scores = model.train(
stimulus=stimulus[:-1],
response=response[:-1],
fs=fs,
tmin=0.0,
tmax=kernel.shape[0] / fs,
regularization=np.logspace(-6, 0, 7),
segment_duration=1.0,
overlap=0.5,
window="hann",
k="loo",
trial_weights=inverse_variance_weights(response[:-1]),
)
prediction = model.predict(stimulus=stimulus[-1])
score = model.score(stimulus=stimulus[-1], response=response[-1])
fig, ax = model.plot(
input_index=0,
output_index=0,
title="Recovered kernel from the held-out example",
)
The plot below is generated from the exact seeded example above:
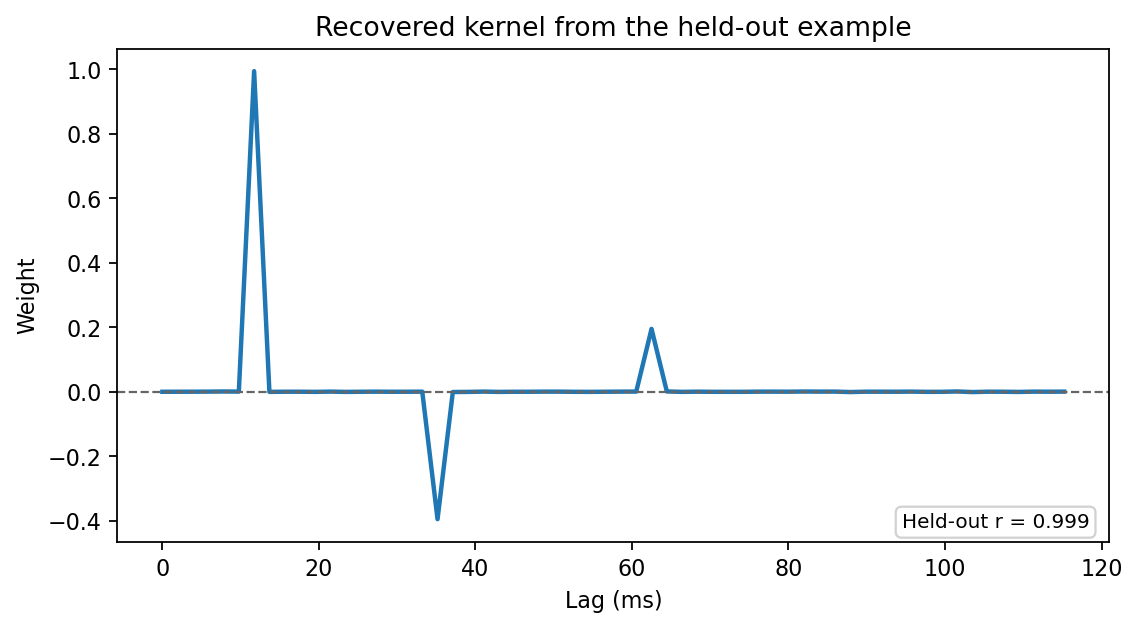
What This Example Is Doing
stimulusandresponseare lists of arrays, so the fit runs in multi-trial mode.stimulus[:-1]andresponse[:-1]are the training trials, while the last trial is held out for evaluation.direction=1means a forward model: stimulus features predict response channels.tmin=0.0andtmax=kernel.shape[0] / fsrequest a causal lag window that matches the simulated ground-truth kernel.regularization=np.logspace(-6, 0, 7)asks the model to cross-validate over seven ridge values instead of fitting one fixed value.segment_duration=1.0sets the segment size used to estimate the cross-spectra.overlap=0.5reuses half of each segment in the next one, which can help stabilize estimates when segments are short.window="hann"applies a Hann window before the FFT.k="loo"means leave-one-out cross-validation over trials.trial_weights=inverse_variance_weights(response[:-1])downweights noisier training trials in the aggregate spectra.prediction = model.predict(...)returns the held-out prediction, andscore = model.score(...)evaluates that held-out trial with the estimator's configured metric.
What train(...) Returns
- If
regularizationis a single scalar,train(...)returnsNoneand fits directly. - If
regularizationcontains multiple candidates,train(...)returns the cross-validation scores for those candidates. - Regardless of return value, the fitted state is stored on the estimator.
After training, the most important attributes are:
model.transfer_function: complex frequency-domain solutionmodel.frequencies: frequency axis in Hzmodel.weights: lag-domain kernelmodel.times: lag axis in secondsmodel.regularization: chosen ridge value or banded tuplemodel.regularization_candidates: all evaluated regularization candidates
What predict(...) Returns
- If you pass only the predictor side,
predict(...)returns predictions. - If you also pass the observed target side, it returns
(predictions, score). - The default score is Pearson correlation, but you can choose a different metric when constructing the model.
score(...)always requires the observed target side and returns only the metric.
Input Conventions
- Single-trial input can be a 1D or 2D NumPy array.
- Multi-trial input should be a list of arrays.
- Each trial must be shaped
(n_samples, n_features)for the predictor side and(n_samples, n_outputs)for the target side. - A 1D vector is treated as a single feature or output channel.
- All stimulus/response pairs within one fit must have matching sample counts per trial.
- In a backward model (
direction=-1), the predictor side is the response and the target side is the stimulus, but the same shape rules apply.
A Minimal Direct Fit
If you already know the ridge value you want, use a direct fit:
model = TRF(direction=1)
model.train(
stimulus=stimulus,
response=response,
fs=fs,
tmin=0.0,
tmax=0.120,
regularization=1e-3,
)
This avoids cross-validation and fits immediately with one regularization setting.
Common Next Actions After Training
model.plot(...): show one kernelmodel.plot_grid(...): show all kernels in a feature-by-output gridmodel.to_impulse_response(...): extract a different lag window without refittingmodel.score(...): compute only the metricmodel.transfer_function_components_at(...): inspect magnitude, phase, and group delaymodel.cross_spectral_diagnostics(...): compare predicted and observed spectramodel.permutation_test(...): compare a held-out score against surrogate null scoresmodel.refit_permutation_test(...): rerun the training pipeline on surrogate-aligned training data for a stronger nullmodel.bootstrap_confidence_interval(...): estimate uncertainty over trialsmodel.save(...): persist the fitted model
Next Steps
- Walk through the intended lifecycle in Core Workflow
- Learn the array rules in Inputs and Shapes
- Learn how regularization search works in Regularization and CV
- Learn how to choose segment length and windowing in Choosing Segment Settings
- See the multitaper workflow in Multitaper Estimation
- Explore time-frequency views in Frequency-Resolved Analysis
- Explore spectral inspection tools in Diagnostics and Transfer Functions
- Learn the held-out significance workflow in Significance Testing
- Browse runnable scripts in Examples
- Open the rendered tutorial in Getting Started Notebook