ffTRF
ffTRF is a Python toolbox for fitting temporal response functions in the
frequency domain. It is designed for continuous stimulus-response modeling
without tying the workflow to one modality, one preprocessing stack, or one
experimental paradigm.
The main estimator is fftrf.TRF. It supports:
- forward and backward TRF fitting
- scalar ridge and cross-validated ridge selection
- optional banded regularization for grouped predictors
- optional DPSS multi-taper spectral estimation
- multi-trial input as Python lists of arrays
- time-domain kernel export for interpretation
- bootstrap confidence intervals
- permutation-based held-out score significance testing
- optional stronger refit-based null-model significance testing
- transfer-function and cross-spectral diagnostics
- frequency-resolved lag-domain views of recovered kernels
Workflow at a Glance
The typical workflow is:
- prepare stimulus and response arrays with matching sample counts per trial
- create
TRF(direction=1)for forward encoding orTRF(direction=-1)for backward decoding - call
train(...)ortrain_multitaper(...) - inspect the fitted kernel with
plot(...)orplot_grid(...) - evaluate generalization with
predict(...),score(...), orpermutation_test(...) - inspect spectral behavior with transfer-function and coherence diagnostics
- save the fitted model with
save(...)if you want to reuse it later
ffTRF stores both the complex frequency-domain transfer function and the
derived lag-domain kernel, so users can move between time-domain and
frequency-domain views without retraining.
Start Here
- New to the package: go to Getting Started
- Prefer a tutorial you can read like a lab notebook: go to Getting Started Notebook
- Need the full workflow explained end to end: go to Core Workflow
- Need shape conventions and multi-trial rules: go to Inputs and Shapes
- Looking for runnable demos: go to Examples
- Looking for API docs: go to Reference
- Working on the repo itself: go to Development
Installation
For local development, Pixi is the primary supported workflow:
pixi install
pixi run import-check
pixi run -e test test
For a lightweight editable install:
pip install -e .
Optional extras:
pip install -e ".[test]"
pip install -e ".[compare]"
pip install -e ".[docs]"
For docs builds, prefer the lockfile-backed Pixi command:
pixi run -e docs docs-build
What ffTRF Actually Fits
The estimator solves for a complex transfer function in the frequency domain
and then transforms that solution into a time-domain kernel over the lag window
you request with tmin and tmax.
That has a few practical consequences:
- you do not need to build an explicit lag matrix yourself
- fitting can reuse cached per-trial spectra across cross-validation folds
- the same fitted model can be inspected in both lag-domain and frequency-domain forms
- optional multi-taper estimation fits into the same API instead of requiring a separate estimator class
Documentation Map
- Getting Started: first fit, first prediction, first plot
- Core Workflow: how the estimator is meant to be used step by step
- Inputs and Shapes: single-trial vs multi-trial conventions and direction-dependent behavior
- Regularization and CV: direct fits, CV grids, and banded ridge
- Choosing Segment Settings: practical
defaults for
segment_duration, overlap, and windowing - Multitaper Estimation: when and how to use DPSS
- Frequency-Resolved Analysis: lag-frequency views of the fitted kernel
- Diagnostics and Transfer Functions: spectral inspection tools
- Trial Weighting and Bootstrap: weighting noisy trials and quantifying uncertainty
- Significance Testing: permutation-based null distributions for held-out scores
- Notebooks: rendered walk-throughs with the same API used in the example scripts
- Reference: detailed function-by-function API
Example Gallery
Single-Trial Forward Model
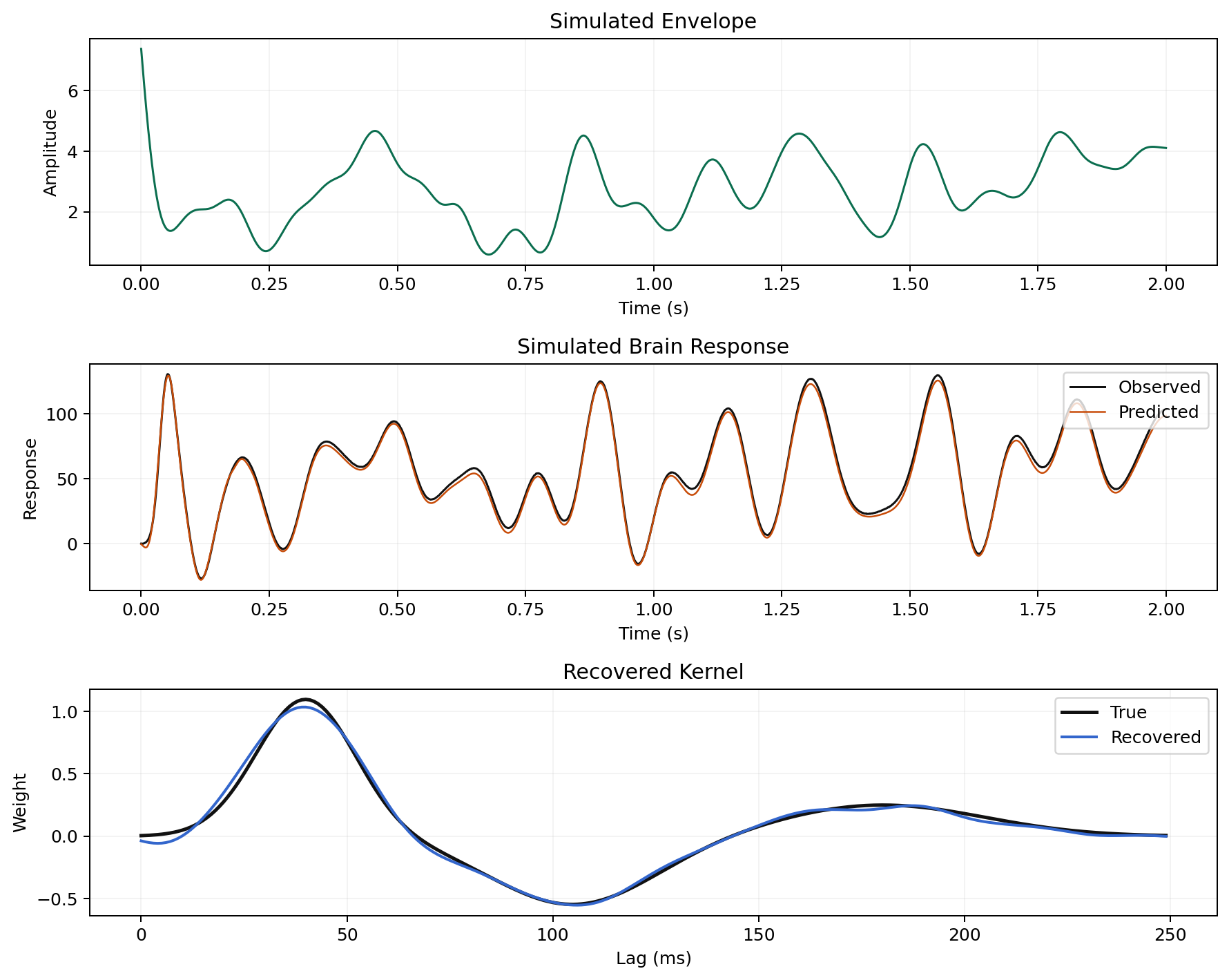
Multi-Trial Cross-Validation
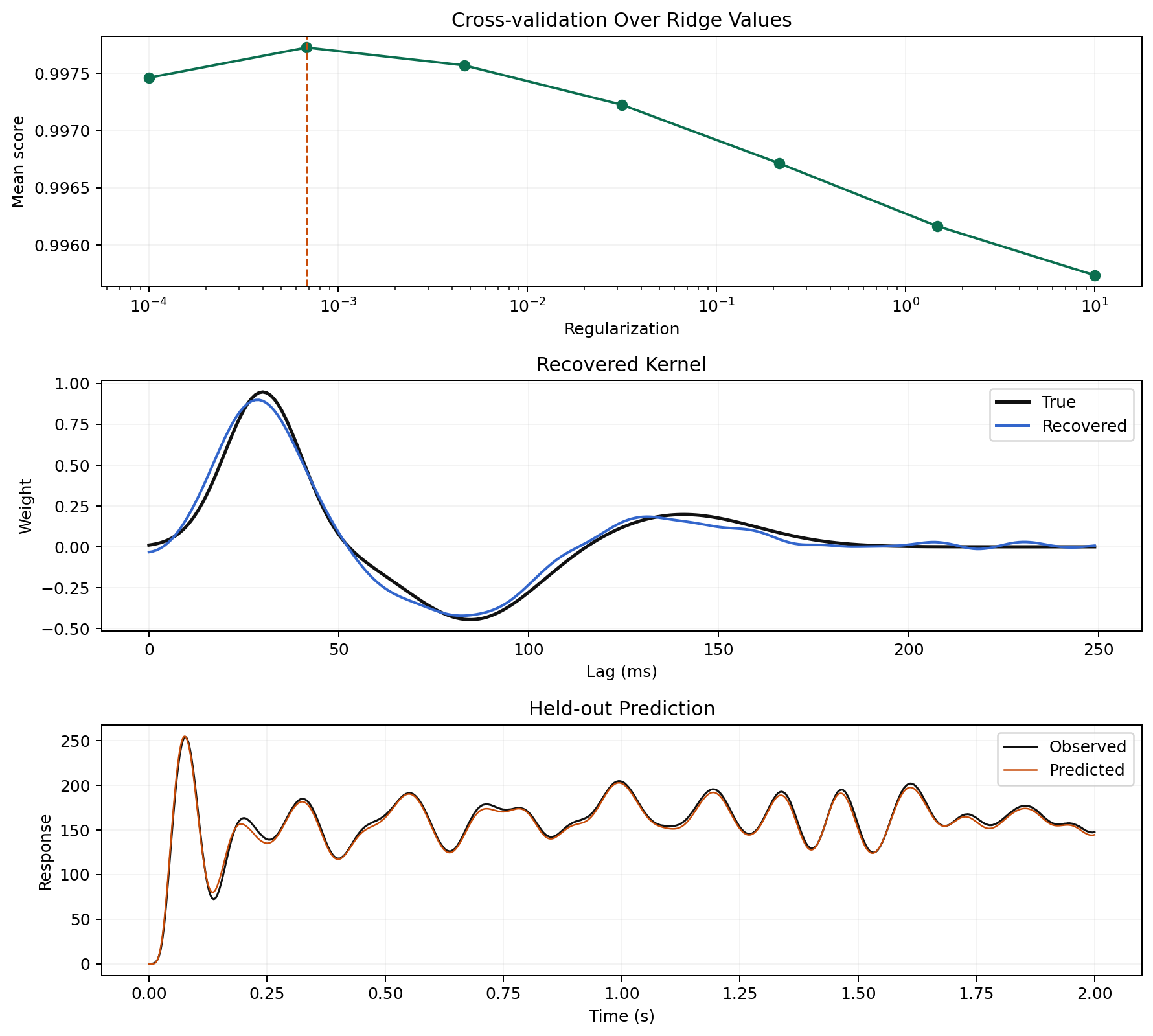
Frequency-Resolved Weights
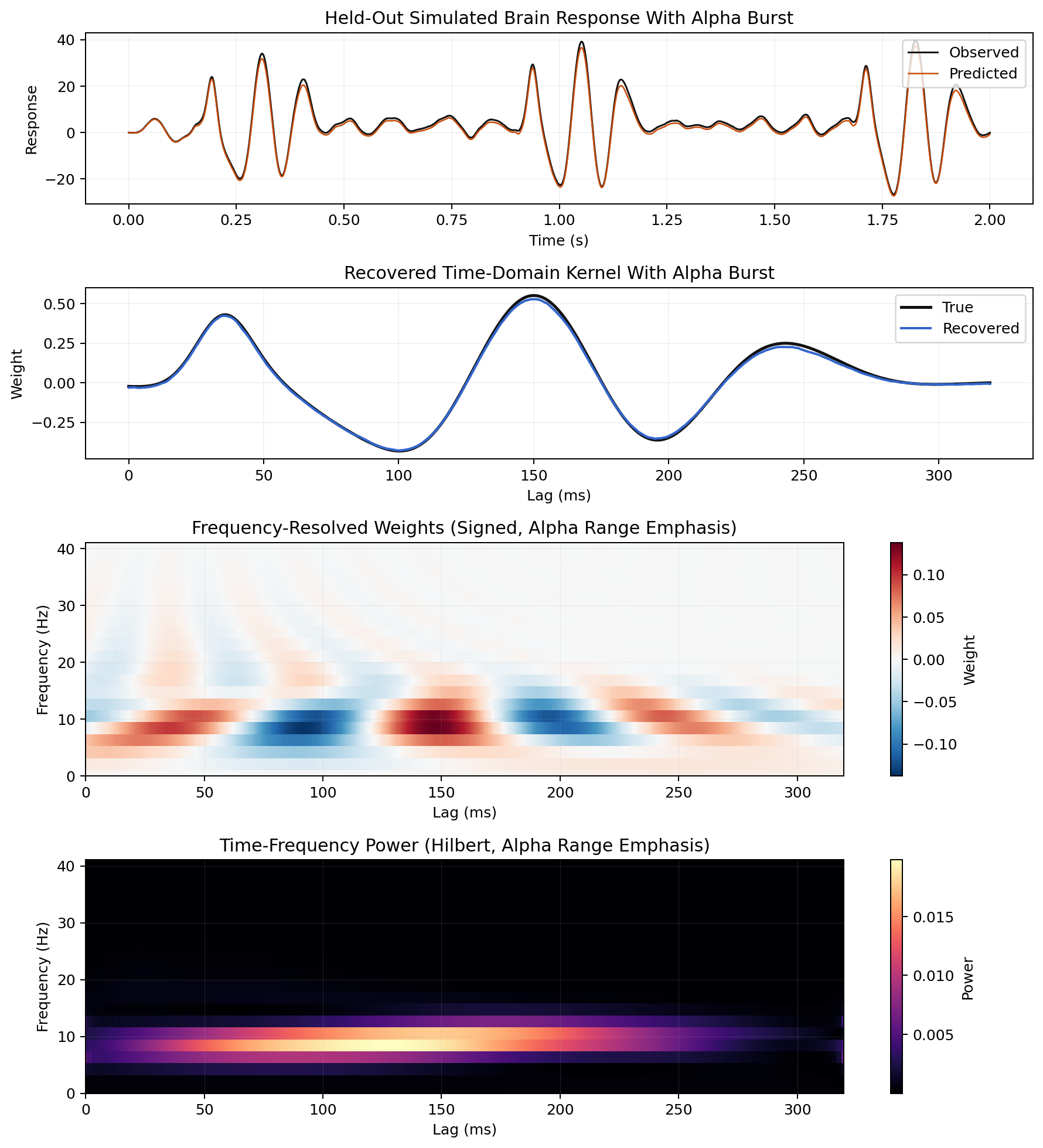
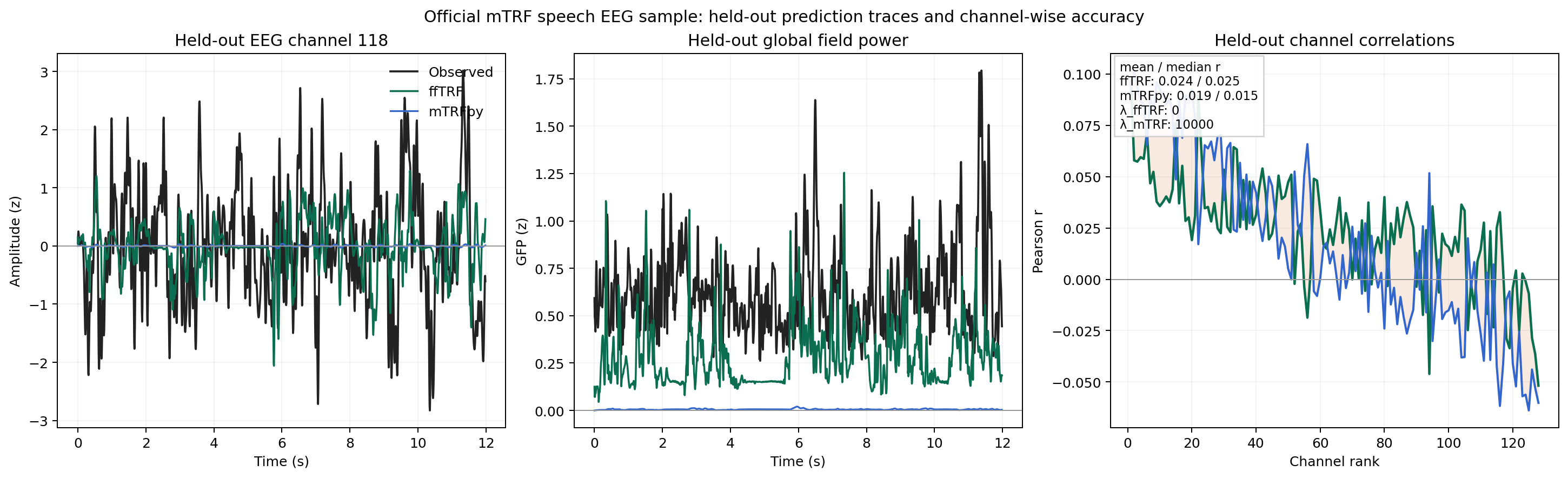
Real EEG Comparison